Author: Cameron Ashby
Date: January 27, 2026
Series: Capstone Development Journal #1
Introduction
I’m Cameron Ashby, a graduate student in the MS Computer Science program at Full Sail University with a concentration in Artificial Intelligence. This fall, I’ll be starting the Doctor of Technology program at Purdue University, focusing on AI transparency and computational biology.
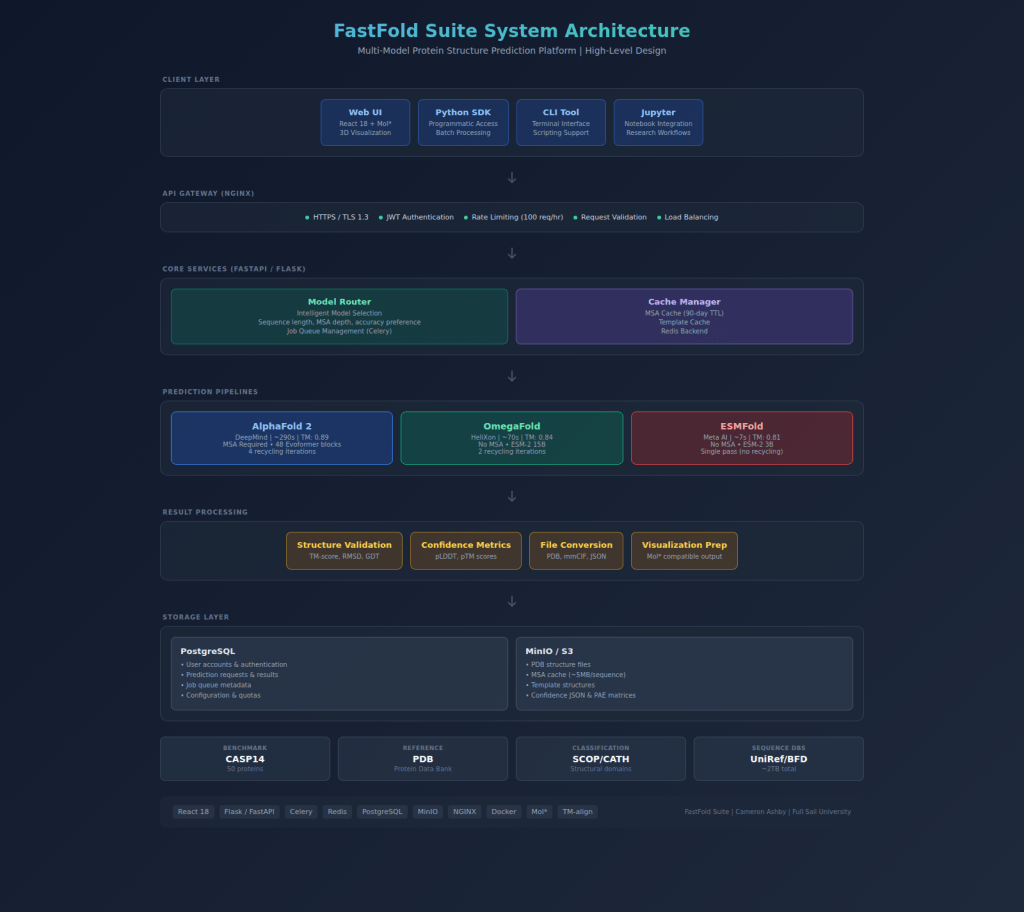
This blog series documents the development of FastFold Suite, my capstone project. FastFold is a web-based platform that integrates multiple AI models for protein structure prediction, specifically ESMFold, OmegaFold, and AlphaFold 2. The goal is to make these powerful tools accessible to researchers who lack the infrastructure or technical background to run them independently.
My background spans broadcast operations (Fox News, CBS, Big Ten Network), educational technology, and now AI/ML development. I approach software the same way I approach live television: build systems where failure isn’t an option, document everything, and make complex workflows accessible to end users.
Month 1: Foundation and Integration Planning
The Problem
AlphaFold changed everything in computational biology. DeepMind’s model achieved near-experimental accuracy in predicting protein structures, a problem scientists had worked on for 50 years. But there’s a gap between “AlphaFold exists” and “I can actually use it.”
Running AlphaFold requires:
- 2TB of database downloads
- CUDA-compatible GPUs
- Linux command-line expertise
- 2-3 weeks of setup time
Most researchers don’t have this infrastructure. Alternatives like ESMFold and OmegaFold are faster and lighter, but they also lack user-friendly interfaces. And there’s no unified way to compare predictions across models.
FastFold Suite solves this.
What I Built (Phase 1 Recap)
From May through October 2025, I completed the HCI capstone phase:
- Roughly 8,000 lines of code across React frontend and Flask backend
- Demo mode with working sequence input, job tracking, and 3D visualization using Mol*
- WCAG 2.1 AA accessibility compliance
- REST API architecture ready for model integration
[IMAGE 1: FastFold Suite interface showing sequence input and 3D protein visualization]
Month 1 Progress (ML Capstone)
This month focused on three areas:
1. AlphaFold 2 Weights Acquisition
I successfully obtained AlphaFold 2 model weights from DeepMind. This was a significant milestone because it expanded my project from a two-model comparison (ESMFold vs. OmegaFold) to a three-model analysis. The research hypothesis was updated accordingly:
“Protein sequence length and structural classification significantly predict the accuracy difference between AlphaFold 2, ESMFold, and OmegaFold predictions.”
2. Capstone Presentation Development
I built a 20-slide presentation covering:
- Protein folding fundamentals for non-experts
- The AlphaFold breakthrough and why it matters
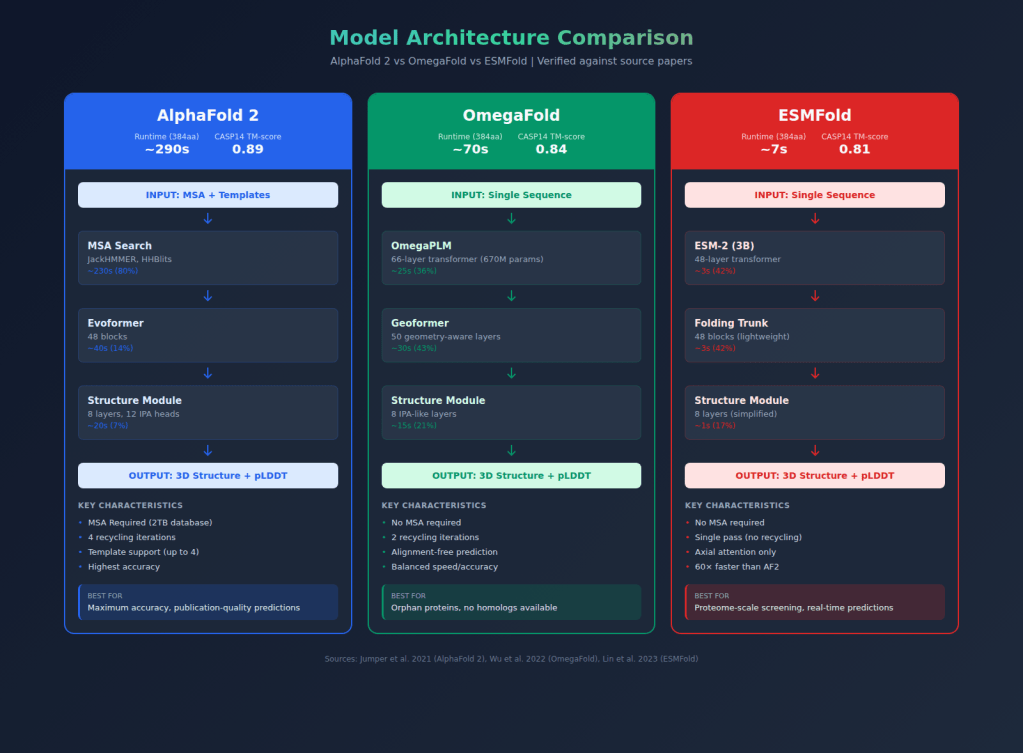
- Technical architecture of all three models
- My benchmarking methodology (50 CASP14 proteins, TM-score/RMSD/pLDDT metrics)
- Statistical analysis plan (Pearson correlation, one-way ANOVA)
3. Documentation and IRB
Completed IRB documentation (exemption request, since this is computational benchmarking with no human subjects) and maintained weekly advisor check-ins with Dr. Andreas Marpaung.
Tools and Resources
- Frontend: React 18, Mol* (3D molecular visualization)
- Backend: Flask, Python
- AI Models: ESMFold (Meta), OmegaFold, AlphaFold 2 (DeepMind)
- Datasets: CASP14, Protein Data Bank (PDB), SCOP/CATH classifications
- References: Jumper et al. 2021 (AlphaFold), Lin et al. 2023 (ESMFold), Wu et al. 2022 (OmegaFold)

Retrospective
What Went Right
- AlphaFold weights acquisition – Getting access earlier than expected gives me more time for integration and testing
- Presentation preparation – Forced me to articulate the technical details clearly, which will help when writing the IEEE paper
- Advisor communication – Weekly check-ins with Dr. Marpaung kept the project on track; added a Saturday session before the presentation for final prep
What Went Wrong
- Scope creep risk – Adding AlphaFold 2 as a third model increases complexity. I need to stay disciplined about the timeline.
- Local GPU limitations – Attempted to run ESMFold locally on Mac M3 Pro, hit CUDA dependency issues. Pivoted to an API-based approach, which worked but required rearchitecting some backend code.
How I’ll Improve
- Stick to the schedule – ESMFold integration must be complete by the end of Week 4. No exceptions.
- Document as I go – The IEEE paper will be easier if I capture methodology details now rather than reconstructing them in April.
- Test incrementally – Validate each model integration against known PDB structures before moving to the next.
Looking Ahead
Month 2 priorities:
- Complete ESMFold integration with end-to-end predictions
- Begin OmegaFold integration
- Build the side-by-side comparison interface
- Implement accuracy metrics (TM-score, RMSD, pLDDT)
The foundation is solid. Now it’s time to make it work.
Contact: cdashby@student.fullsail.edu
GitHub: https://github.com/cashby70
LinkedIn: https://www.linkedin.com/in/cameron-ashby-mfa-8217a7241/
This post is part of my MS Computer Science capstone journal at Full Sail University.
Leave a comment